Details
-
Type:
Task
-
Status: Closed (View Workflow)
-
Priority:
Major
-
Resolution: Done
-
Affects Version/s: None
-
Fix Version/s: None
-
Labels:None
-
Story Points:1
-
Sprint:Fall 4 2023 Oct 16, Fall 5
Description
File: FindTreatmentEffectAcrossGenotypes-DESeq2.Rmd
Check to see if the analysis and the interaction term is explained well and provides enough sanity checking to confirm no significance.
Use bar plot shiny app to show analysis.
During meeting with Muday Lab, we found a gene that we thought ought to have come up in a test of the interaction term: Solyc02g063355.1
The SL5 version of that gene appears in the top row of the attached heat map file 2023-10-19-meeting-heatmap-visualization.png.
The heat map shows log2(average(cpm(A.28/V.28))) for treatment durations indicated on the x axis of the plot.
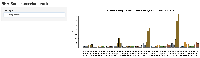



Confirming the Analysis with bar plots of gene expression: