Details
-
Type:
Documentation
-
Status: Closed (View Workflow)
-
Priority:
Major
-
Resolution: Done
-
Affects Version/s: None
-
Fix Version/s: None
-
Labels:None
-
Story Points:1
-
Epic Link:
-
Sprint:Summer 1, Summer 2, Summer 3
Description
In the most recent IGB workshop, a participant asked if they could view two genomes at once in IGB.
In the News section on the BioViz Web site (see: https://bioviz.org/news.html) we wrote:
Q: Can I load two genomes at once in IGB?
A: IGB does not support viewing two genomes at once. However, you can get around this by opening another IGB window to view a second genome. Contact us if you need help.
I think that if you run IGB "from the command line," you can launch more than one IGB process and get one IGB window per process.
For this task, check into whether we already have documentation about this in our user's guide.
If we do not, then create documentation explaining how to do this, and provide a link of some type from the News item to the User's Guide.
Attachments
Issue Links
- relates to
-
IGBF-4185 Add BiAS Workshop to IGB News
-
- Closed
-
Activity
| Field | Original Value | New Value |
|---|---|---|
| Epic Link | IGBF-140 [ 14594 ] |
| Sprint | Spring 7 [ 216 ] | Spring 8 [ 217 ] |
| Status | To-Do [ 10305 ] | In Progress [ 3 ] |
| Status | In Progress [ 3 ] | Needs 1st Level Review [ 10005 ] |
| Assignee | Paige Kulzer [ pkulzer ] | Nowlan Freese [ nfreese ] |
| Sprint | Spring 8 [ 217 ] | Summer 1 [ 218 ] |
| Attachment | SL4-ARE-zoomed-out-and-selected.png [ 18697 ] | |
| Attachment | SL5-ARE-zoomed-out-and-selected.png [ 18698 ] |
| Attachment | SL4-ARE-zoomed-out-and-selected.png [ 18697 ] |
| Attachment | SL4-ARE-zoomed-out-and-selected.png [ 18699 ] |
| Attachment | SL4-ARE-alignments-S1.png [ 18700 ] | |
| Attachment | SL4-ARE-alignments-S3.png [ 18701 ] |
| Attachment | SL4-ARE-S1-alignments-selected-and-counted.png [ 18702 ] | |
| Attachment | SL4-ARE-S3-alignments-selected-and-counted.png [ 18703 ] |
| Sprint | Summer 1 [ 218 ] | Summer 1, Summer 2 [ 218, 219 ] |
| Rank | Ranked higher |
| Sprint | Summer 1, Summer 2 [ 218, 219 ] | Summer 1, Summer 2, Summer 3 [ 218, 219, 220 ] |
| Rank | Ranked higher |
| Comment |
[ FYI:
I am uploading the alignments to RENCI host in preparation for adding the data to an SL4 Quickload. ] |
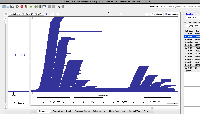
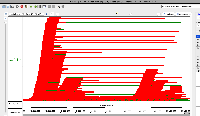
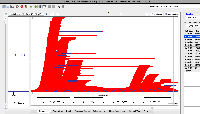
| Comment | [ I hypothesized that the reason I saw more enormous-intron possibly spurious alignments in S1 was that S1 simply has more alignments overall. To test this, I click-dragged over all the alignments in each view to select them. IGB then reported the number of selected items in the Selection Info text box in the top right of the window. The resulting images are attached. ] |
| Comment | [ Two Igb's are better than one. ] |
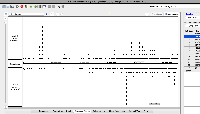
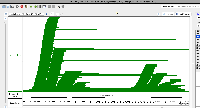
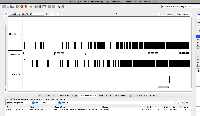
| Comment | [ Another use case for multi-window IGB visualization is shown in the next two attached files. Both show the same locus on the same genome assembly with data from two single cell RNA-Seq data sets loaded. One shows what appears to be more spurious very-long-intron alignments relative to the entire data set. However, it is not clear from the images which data set contains more alignments in total. ] |
| Comment | [ I attach two example IGB window images. Each show the location of the same gene but in a different assembly of S. lycopersicum. Check the top of the IGB window for more information about the assemblies being shown, such as synonyms. ] |
| Status | Needs 1st Level Review [ 10005 ] | First Level Review in Progress [ 10301 ] |
| Status | First Level Review in Progress [ 10301 ] | Ready for Pull Request [ 10304 ] |
| Status | Ready for Pull Request [ 10304 ] | Pull Request Submitted [ 10101 ] |
| Status | Pull Request Submitted [ 10101 ] | Reviewing Pull Request [ 10303 ] |
| Status | Reviewing Pull Request [ 10303 ] | Merged Needs Testing [ 10002 ] |
| Status | Merged Needs Testing [ 10002 ] | Post-merge Testing In Progress [ 10003 ] |
| Resolution | Done [ 10000 ] | |
| Status | Post-merge Testing In Progress [ 10003 ] | Closed [ 6 ] |