Details
-
Type:
Bug
-
Status: Closed (View Workflow)
-
Priority:
Major
-
Resolution: Done
-
Affects Version/s: None
-
Fix Version/s: 8.5.2 MInor Release
-
Labels:
-
Story Points:0.25
-
Sprint:Sprint 30
Description
Insertion Glyph bases are not being shown correctly.
The Glyph is black (correct) but the bases are no longer being shown.
To repeat:
1) Visit S. lycopersicum (tomato) genome
2) Select chromosome chrC-NC_007898.3
3) Select a dataset from Plastid Resequencing folder (not Primers or Amplicon - choose one of the others)
4) Zoom in, load data
5) Zoom in further, look for black insertion Glyphs
6) Observe that bases are not being shown.
What should happen is that bases flanking the insertion site should be displayed on top of the black glyph.
Attachments
Issue Links
- is duplicated by
-
IGBF-885 Change Insertion glyph color and shape
-
- Closed
-


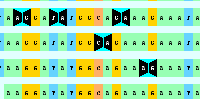

While testing this issue I noticed that VCF insertions only show one base pair letter on the insertion rather than 2 (see screenshot "VCF_Insertion.png").
I am waiting on feedback to determine if this is expected or not, but if it is then this story can be closed since all other aspects of this story have passed testing.