Details
-
Type:
Bug
-
Status: Closed (View Workflow)
-
Priority:
Major
-
Resolution: Done
-
Affects Version/s: None
-
Fix Version/s: None
-
Labels:None
-
Story Points:4
-
Epic Link:
-
Sprint:Spring 3 : 17 Feb to 28 Feb, Spring 4 : 2 Mar to 13 Mar, Spring 6 : 30 Mar to Apr 10, Spring 7 : 13 Apr to 24 Apr, Spring 8 : 24 Apr to 8 May, Spring 8 : 11 May to 25 May, Spring 9 : 25 May to 8 Jun, Summer 1: 8 Jun - 19 Jun
Description
When we first started migrating to OSGi, developer Lance Frohman (Genentech) created a plug-in that drew a sashimi-plot graphic on top of gene models to highlight splicing support.
Recently, we moved the code to a new repository:
https://bitbucket.org/lorainelab/development-apps/src/master/TranscriptIsoform/
Let's revisit this code and investigate what it would be take to implement it now.
Attachments
Issue Links
- blocks
-
IGBF-2373 Move transcript isoform App into a separate repository
-
- Closed
-
Activity
Prof. [~aloraine], please correct me if I am wrong.
I have tried a few ways to add submenu but I am a bit confused about things mentioned below:
- When we create a set of MenuItem to add as a submenu and when it is passed to a function MenuItem(String menuLabel, Set<MenuItem> subMenuItems) of MenuItem, this Set is assigned to a private "subMenuItems" property of the class and in order to add to to the actual menu in IGB it should be accessed with the help of Set<MenuItem> getSubMenuItems() function of the class.
-When I tried searching for the call to "getSubMenuItems" function mentioned above in the IGB codebase, I only found it in the "SeqMapView" class file which is used to add submenu items in the "popup menu" which I think is called upon the selection of Syms. In that code it iterates over the list Set of MenuItems receives with the help of Set<MenuItem> getSubMenuItems() function of MenuItem class. - According to me, there should be a code in IGB which is triggered when we add the new MenuItem in IGB Menu which iterates over the list Set of MenuItems receives with the help of Set<MenuItem> getSubMenuItems() function of MenuItem class as well. I couldn't find a piece of code which manipulates IGB menu by calling "getSubMenuItems" Set which provides a list of MenuItems that should be added in submenu.
- We might need to change a few things on IGB side for that.
Also, I referred the plugins which are available in IGB code as well as the apps which are available and found that no other app is trying to add a submenu in IGB dynamically.
Dr. Loraine, if you know any old app which tries to add menu with a submenu in IGB at run time then please let me know the name of that app.
It would be great if you could suggest something in this situation.
- To check out the difference between (1) IgbMenuItemProvider and (2) MenuBarEntryProvider refer:
(1) https://bitbucket.org/lorainelab/integrated-genome-browser/src/igb_8_4/core/igb-services-api/src/main/java/com/lorainelab/igb/services/window/menus/IgbMenuItemProvider.java
(2) https://bitbucket.org/lorainelab/integrated-genome-browser/src/igb-9.0.0-pre-rollback/core/menu-api/src/main/java/org/lorainelab/igb/menu/api/MenuBarEntryProvider.java
- Also, I was working on IGB 8.2 to check the functionality of the older version of this app.
- I checked out to "igb_8_2" branch, then I changed a few things in the pom to make the plugin work. The old plugin had packaging type JAR.
- I changed it to bundle in order to generate repository.xml
- POM changes:
<project xmlns="http://maven.apache.org/POM/4.0.0" xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance" xsi:schemaLocation="http://maven.apache.org/POM/4.0.0 http://maven.apache.org/xsd/maven-4.0.0.xsd"> <modelVersion>4.0.0</modelVersion> <parent> <groupId>com.affymetrix</groupId> <artifactId>igb-project</artifactId> <version>8.2</version> <relativePath>../../pom.xml</relativePath> </parent> <groupId>com.affymetrix</groupId> <artifactId>transcriptIsoForm</artifactId> <version>0.0.1</version> <packaging>bundle</packaging> <name>transcriptIsoForm</name> <dependencies> <dependency> <groupId>org.apache.felix</groupId> <artifactId>org.apache.felix.main</artifactId> <scope>provided</scope> </dependency> <dependency> <groupId>com.affymetrix</groupId> <artifactId>igb</artifactId> <scope>provided</scope> </dependency> <dependency> <groupId>net.sf</groupId> <artifactId>sam-ext</artifactId> <scope>provided</scope> </dependency> <dependency> <groupId>com.affymetrix</groupId> <artifactId>genoviz</artifactId> <scope>provided</scope> </dependency> <dependency> <groupId>com.affymetrix</groupId> <artifactId>igb-service</artifactId> <scope>provided</scope> </dependency> <dependency> <groupId>com.affymetrix</groupId> <artifactId>igbSwingExt</artifactId> <scope>provided</scope> </dependency> <dependency> <groupId>net.sf</groupId> <artifactId>igb-picard</artifactId> <scope>provided</scope> </dependency> <dependency> <groupId>com.affymetrix</groupId> <artifactId>genometryImpl</artifactId> <scope>provided</scope> </dependency> </dependencies> <build> <plugins> <plugin> <groupId>org.apache.felix</groupId> <artifactId>maven-bundle-plugin</artifactId> <extensions>true</extensions> <executions> <execution> <id>build plugin repository</id> <goals> <goal> index </goal> </goals> <phase>package</phase> <configuration> <obrRepository>${project.build.directory}</obrRepository> <mavenRepository>${project.build.directory}</mavenRepository> </configuration> </execution> <execution> <id>build obr plugin repository</id> <goals> <goal> bundle </goal> </goals> <configuration> <instructions> <Include-Resource>${project.build.directory}/repository.xml, {maven-resources}</Include-Resource> </instructions> </configuration> </execution> </executions> <configuration> <instructions> <Bundle-SymbolicName>${project.artifactId}</Bundle-SymbolicName> <Export-Package/> <Bundle-Activator>com.gene.transcriptisoform.Activator</Bundle-Activator> <Bundle-ActivationPolicy>lazy</Bundle-ActivationPolicy> <Bundle-Description>${bundleDescription}</Bundle-Description> <Service-Component>*</Service-Component> </instructions> </configuration> </plugin> <plugin> <groupId>org.lorainelab</groupId> <artifactId>bundle-markdown-encoder</artifactId> <version>1.0.0</version> <executions> <execution> <goals> <goal>encodeMarkdown</goal> </goals> </execution> </executions> </plugin> </plugins> </build> </project>
- The plugin was installed successfully after making these changes. Attaching a screenshot of the menu to the ticket for future reference.
- Then as discussed, I tried to figure out the functionality of the app by selecting RefSeq track. I thought a graph will be shown but nothing appeared on the screen. Even when I changed the brightness and thickness options from the "View" menu.
- The older code is using Activator class and for displaying menu with submenus it is using "com.affymetrix.genoviz.swing.AMenuItem".
Progress:
- I tried to replicate the old functionality by pasting "Activator" class in the old code directly to the newer one just to make sure the menu is getting rendered properly.
- Then I changed small sections of it, to make it like the newer one by removing ServiceRegistrar and with the reference of NCBIPrimer older and newer version as event NCBI app used to have an "Activator" class.
- I am still facing a few issues with it which I am going to work on today.
Progress:
- I am able to add the submenu and submenu to the submenus.
- We have also added the actionListeners to these menu items today.
- We are not sure how these options work.
Prof. [~aloraine], do you have any idea about it?
I have shared the IGB 8.2 code with you just now which has a working old version of the app inside developmentPlugins directory.
Could you please install it and let us know what exactly those submenu items do?
It would be great if you mention the steps to check the functionality.
cc: Shamika Gajanan Kulkarni
Thanks for the zip file! It was fun to run an older version of IGB.
Workflow to demo Transcript Isoform App
- Open Arabidopsis thaliana genome
- Under Data Access panel, open an RNA-Seq data set. For examle, if you are running this older version of IGB, open Pollen > RNA-Seq > SM > Reads > Pollen, Alignments.
- Zoom in on a gene that has some introns. Click Load Data to load some read alignments into the browser. If you are using this older version of IGB, use the Advanced Search tab Keyword Search to find genes that have "pollen" in the description. This will ensure you find some genes that are actually expressed in pollen.
- Select a reference track: Choose View > Transcript Isoform > Select Reference Tiers.
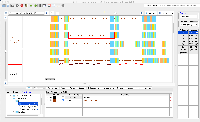
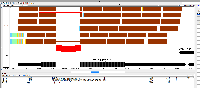
- What should happen next is that a "chevron" (looks like a "v") should appear above introns in the reference tier wherever there are "gapped" reads that span the intron.
This chevron graphic is meant to visually summarize the number of read alignments that support each intron shown in the reference tier.
Question:
- How did you get Jira to insert the pom.xml code?
See attached image for Transcript Isoform running in IGB 8.2.0.
Pro. [~aloraine]
To insert POM I enclosed the code inside
{code:xml}<insert you code here>{code}
I thought it would be convenient to check the code in the comment that way as it is shown in a different window which preserved the formatting.
Like xml there are different tags which we can use which can be explored using the link present below:
https://confluence.atlassian.com/doc/code-block-macro-139390.html
Changes are present at:
https://bitbucket.org/pkulka10/experimental-apps-for-igb/commits/51a3f4f09268f72eaf395f9ff36b9bff16cc8171
Please follow the steps "Workflow to demo Transcript Isoform App" as mentioned in the previous comment.
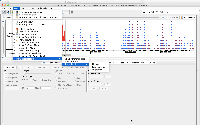
Also, I was trying to run it one last time with latest master's mac installer using the same workflow but IGB's quickload was throwing some exception which is attached as an image to the ticket.
I think it is happening because of large amount of data.
Yes, the user may need to zoom in to a region and only then select the reference tier. This is obviously not the best design. We will need to better understand how to throttle loading of the alignments.
However, it looks like you have got it working - a good first draft.
There is more work to be done on this, but not today! Do please submit PR.
Prutha Kulkarni
(Note that the next step will be moving the code, with history intact, into a new repository.)
Can you do this one next - after the PR is merged?
Sure. I will do that asap
PR submitted.
code changes can be found at:
https://bitbucket.org/lorainelab/experimental-apps-for-igb/pull-requests/12/igbf-2123-refactor-transcriptisoform-app/diff
PR merged.
- commit:
IGBF-2123: Refactor TranscriptIsoform App
is now on master branch.
cc: Prutha Kulkarni
Note this cannot be tested until the bitbucket pipelines are configured properly at the team repository at https://bitbucket.org/lorainelab/transcriptisoform. Hence IGBF-2373 is a blocker for testing.
Tested on the latest dev version of IGB on macOS.
- The expected behavior described in this comment occurs.
- Toggling between the View > Transcript Isoform > Show density options results in visually different 'chevrons' in the intergenic reference tiers. I am not sure of the significance of these differences or what they are expected to be, but the functionality of the app from the new repo seems to resemble that of the same app from the experimental apps repo.
- I noticed that the build at the new repo fails, but since
IGBF-2373has been resolved, it may be good to try the build again. - Some intergenic regions have multiple overlapping 'chevrons' on the reference tiers. Is this expected? I have attached a screenshot (overlapping_chevrons.png) - the same behavior seems to be present when running the app from the experimental repo.
The app seems to be functioning as expected. Moving to done.

Progress:
https://bitbucket.org/pkulka10/developmentplugins/commits/e2b1c688e48d5e40e1e8e8665791605fcc2a4ed5#chg-TranscriptIsoform/src/main/java/com/gene/transcriptisoform/TranscriptIsoFormMenuProvider.java
https://bitbucket.org/pkulka10/developmentplugins/commits/f74e0b2065ae033950cdbffbf46e1d32749f255d
MenuItem(String menuLabel, Set<MenuItem> subMenuItems)
as this should be used to add submenu items in IGB menu item.
I have assigned available/unavailable tag next to the types which are provide/not provided in "org.lorainelab.igb.menu.api.model" package.