Details
-
Type:
Improvement
-
Status: Closed (View Workflow)
-
Priority:
Major
-
Resolution: Done
-
Affects Version/s: None
-
Fix Version/s: None
-
Labels:None
-
Story Points:0.25
-
Epic Link:
-
Sprint:Fall 6 2021 Oct 25 - Nov 5
Description
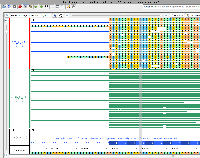
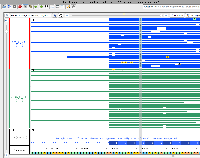
I have made the attached images showing alignment. These need to be added to the repository and documented in the git commit message.
"are" gene: Solyc02g083860
Attachments
Issue Links
- relates to
-
IGBF-2994 Sanity-check expression data by plotting Solyc02g083860 expression
-
- Closed
-
Commit message:
The images help show that the samples labeled "are" are indeed from that genotype. The region shown depicts the version 4 assembly for chromosome 2 and a part of the gene model proposed for the F3H enzyme.
Since all "are" sample reads contain what looks like a point mutation relative to the Heinz reference, this is likely to be the defect described in the literature. According to the image, a C/G base pair (forward strand of chromosome SL4.0ch02, shown in the browser in "Coordinates" track) is converted to T/A in the "are" genotype. The sense strand of the gene is encoded on the reverse strand of the chromosome, and so the browser shows transcription as proceeding right to left, with the top strand serving as the template. The affected codon (5'-AGC-3', the sense strand of the gene) encodes serine in VF36 and Heinz, but is converted to asparagine (5'-AAC-3') in the "are" sample, according to the images shown.