Details
-
Type:
Epic
-
Status: To-Do (View Workflow)
-
Priority:
Major
-
Resolution: Unresolved
-
Affects Version/s: None
-
Fix Version/s: None
-
Labels:None
-
Epic Name:Process and deploy Palanivelu Lab data
-
Story Points:4
-
Sprint:Spring 3 2023 Feb 1, Spring 8 2023 Apr 24, Spring 9 2023 May 1
Description
For this task,
- process new and old experimental data sets from the Ravi Palanivelu lab
- confer with Palanivelu lab personnel to understand and document the samples
- track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/
About the data:
As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to "batches" of RNA samples that were sent to GeneWiz/Azenta for library synthesis and subsequent sequencing.
For two of these batches, the RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta (formerly GeneWiz), the sequencing company. For one of these batches, the so-called "library synthesis pilot," the RP lab synthesized the libraries themselves and then sent the libraries to Azenta.
Once the sequencing data are complete, the company sent links to an ftp site containing the data files to the RP lab, who downloaded them or asked us to download them. We then obtained the sequence data and deployed them to the Charlotte HPC file system for the next steps - data processing, in which we generate files for visualization in IGB and, also, "counts" files for statistical analysis libraries developed for RNA-Seq data.
The three data collections are:
1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536
These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type represented in the sequencing data, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The biological material were created in 2021, in the summer and early spring.
The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the biological material were created separately from the un-pollinated pistils.
Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536.
We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset.
KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the git repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021.
When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings.
2) self-pollinated stigma+style heat stress experiment, Azenta id 30-804059537
We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files.
The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie
This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were:
- temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control)
- treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours
- four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas
- tissue type: dissected stigma and style tissue from self-pollinated flowers
There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples.
Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples
3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment, Azenta id 30-605730043
This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas plants.
The data from this experiment are stored on the UNC Charlotte cluster in: /projects/tomato_genome/rnaseq/ravi-tamaulipas
Rob downloaded these data from the sequencing provider on or around December 15, 2021. (This is the date that Rob created a Google Doc describing the files available from the sequencing provider's data transfer ftp site.)
Note: We need to confirm if that the sequences obtained from the unpollinated pistils were from the same experiment as (1) above. If yes, which "replicate" were they? This will influence how we label the data in IGB.
To-do for each experimental data set:
- Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblies using "reverse" strandedness parameter.
- Check the multi-qc report. Re-run the processing as necessary.
- Rename BAM files to not included "sorted" in the name.
- Create scaled coverage graphs.
- Create junction files.
- Migrate data to an on-line location for IGB visualization.
- Create annots.xml metadata file with visualization parameters for each dataset; add the data collection to the makeAnnotsXml.py script
- Add the "counts" data files to the repository for statistical analysis
- Add documentation for each sequence collection to the git repository
- Perform data checking to catch any record-keeping errors that may have occurred
Attached:
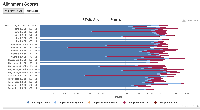
- Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced
- Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing
Contact:
- Kelsey Pryze - kelseypryze@email.arizona.edu
Attachments
Issue Links
Activity
| Field | Original Value | New Value |
|---|---|---|
| Epic Link | IGBF-2993 [ 21429 ] |
| Description | RR to describe location, etc. |
For this task, process new data set from the Palanivelu lab.
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Report from the sequencer (Azenta) is attached. To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on just one sample to get a preliminary MultiQC report. Check the strandedness parameter. * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on all the samples using the correct strandedness parameter. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization |
| Attachment | Azenta_30-804059537_Data_Report.html [ 17672 ] |
| Description |
For this task, process new data set from the Palanivelu lab.
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Report from the sequencer (Azenta) is attached. To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on just one sample to get a preliminary MultiQC report. Check the strandedness parameter. * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on all the samples using the correct strandedness parameter. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization |
For this task, process new data set from the Palanivelu lab.
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Report from the sequencer (Azenta) is attached. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on just one sample to get a preliminary MultiQC report. Check the strandedness parameter. * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on all the samples using the correct strandedness parameter. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization |
| Description |
For this task, process new data set from the Palanivelu lab.
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Report from the sequencer (Azenta) is attached. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on just one sample to get a preliminary MultiQC report. Check the strandedness parameter. * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on all the samples using the correct strandedness parameter. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization |
For this task, process new data set from the Palanivelu lab.
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Report from the sequencer (Azenta) is attached. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on all the samples "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization |
| Attachment | 30-804059537.pdf [ 17673 ] |
| Description |
For this task, process new data set from the Palanivelu lab.
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Report from the sequencer (Azenta) is attached. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on all the samples "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization |
For this task, process new data set from the Palanivelu lab.
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on all the samples "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing |
| Description |
For this task, process new data set from the Palanivelu lab.
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on all the samples "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on all the samples "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing |
| Description |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on all the samples "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie Bitbucket repo: https://bitbucket.org/hotpollen/ovary-rnaseq/src/main/ To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on all the samples "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing |
| Description |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie Bitbucket repo: https://bitbucket.org/hotpollen/ovary-rnaseq/src/main/ To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly on all the samples "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie Bitbucket repo: https://bitbucket.org/hotpollen/ovary-rnaseq/src/main/ To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing |
| Description |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie Bitbucket repo: https://bitbucket.org/hotpollen/ovary-rnaseq/src/main/ To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie Bitbucket repo: https://bitbucket.org/hotpollen/pistil-rna-seq To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing |
| Description |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie Bitbucket repo: https://bitbucket.org/hotpollen/pistil-rna-seq To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie Bitbucket repo: https://bitbucket.org/hotpollen/pistil-rna-seq To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Description |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-kelsie Bitbucket repo: https://bitbucket.org/hotpollen/pistil-rna-seq To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-KP (kp for "Kelsey Pryze") Bitbucket repo: https://bitbucket.org/hotpollen/pistil-rna-seq To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Attachment | KP_samples.csv [ 17674 ] |
| Status | To-Do [ 10305 ] | In Progress [ 3 ] |
| Assignee | Molly Davis [ molly ] |
| Attachment | Screen Shot 2023-02-09 at 9.40.46 AM.png [ 17678 ] |
| Attachment | KP_multiqc_report.html [ 17679 ] |
| Attachment | Screen Shot 2023-02-09 at 10.28.15 AM.png [ 17680 ] |
| Attachment | Screen Shot 2023-02-09 at 10.28.15 AM.png [ 17681 ] |
| Attachment | Screen Shot 2023-02-09 at 10.28.15 AM.png [ 17681 ] |
| Attachment | Screen Shot 2023-02-09 at 10.35.27 AM.png [ 17682 ] |
| Attachment | KP_samples.csv [ 17686 ] |
| Attachment | KP_multiqc_report.html [ 17687 ] |
| Attachment | Screen Shot 2023-02-09 at 10.28.15 AM.png [ 17680 ] |
| Attachment | multiqc_star.txt [ 17688 ] |
| Attachment | KP_samples.csv [ 17674 ] |
| Attachment | KP_multiqc_report.html [ 17679 ] |
| Attachment | Heinz-Ovary-Rep1-0hr-25C-unpol_R1_001_fastqc.html [ 17689 ] |
| Attachment | Heinz-Ovary-Rep2-0hr-25C-unpol_R1_001_fastqc.html [ 17690 ] |
| Attachment | mergedsamtoolsOut.txt [ 17707 ] |
| Status | In Progress [ 3 ] | Needs 1st Level Review [ 10005 ] |
| Description |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-KP (kp for "Kelsey Pryze") Bitbucket repo: https://bitbucket.org/hotpollen/pistil-rna-seq To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attack to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization * Create annots.xml metadata file for visualization Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-KP (kp for "Kelsey Pryze") Bitbucket repo: https://bitbucket.org/hotpollen/pistil-rna-seq To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attach to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters. Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Link | This issue relates to IGBF-3261 [ IGBF-3261 ] |
| Sprint | Spring 3 2023 Feb 1 [ 163 ] | Spring 3 2023 Feb 1, Spring 4 2023 Feb 21 [ 163, 164 ] |
| Rank | Ranked higher |
| Story Points | 2 | 4 |
| Assignee | Molly Davis [ molly ] |
| Status | Needs 1st Level Review [ 10005 ] | First Level Review in Progress [ 10301 ] |
| Status | First Level Review in Progress [ 10301 ] | To-Do [ 10305 ] |
| Assignee | Molly Davis [ molly ] |
| Sprint | Spring 3 2023 Feb 1, Spring 4 2023 Feb 21 [ 163, 164 ] | Spring 3 2023 Feb 1, Spring 5 2023 Mar 6 [ 163, 165 ] |
| Sprint | Spring 3 2023 Feb 1, Spring 5 2023 Mar 6 [ 163, 165 ] | Spring 3 2023 Feb 1, Spring 6 2023 Mar 20 [ 163, 166 ] |
| Sprint | Spring 3 2023 Feb 1, Spring 6 2023 Mar 20 [ 163, 166 ] | Spring 3 2023 Feb 1, Spring 7 2023 Apr 10 [ 163, 167 ] |
| Rank | Ranked higher |
| Sprint | Spring 3 2023 Feb 1, Spring 7 2023 Apr 10 [ 163, 167 ] | Spring 3 2023 Feb 1 [ 163 ] |
| Issue Type | Task [ 3 ] | Epic [ 10000 ] |
| Sprint | Spring 3 2023 Feb 1 [ 163 ] | Spring 3 2023 Feb 1, Spring 8 2023 Apr 24 [ 163, 168 ] |
| Epic Link | IGBF-2993 [ 21429 ] |
| Epic Name | Process Kelsey's Palanivelu Lab data |
| Epic Child |
|
| Epic Child |
|
| Epic Color | ghx-label-6 |
| Epic Child |
|
| Epic Child |
|
| Comment |
[ Update:
* Created sample sheet: [^KP_samples.csv] * Started Nextflow pipeline. ] |
| Epic Child |
|
| Sprint | Spring 3 2023 Feb 1, Spring 8 2023 Apr 24 [ 163, 168 ] | Spring 3 2023 Feb 1, Spring 8 2023 Apr 24, Spring 9 2023 May 8 [ 163, 168, 169 ] |
| Rank | Ranked higher |
| Attachment | Heinz-Ovary-Rep1-0hr-25C-unpol_R1_001_fastqc.html [ 17689 ] |
| Attachment | Heinz-Ovary-Rep2-0hr-25C-unpol_R1_001_fastqc.html [ 17690 ] |
| Status | To-Do [ 10305 ] | In Progress [ 3 ] |
| Status | In Progress [ 3 ] | Needs 1st Level Review [ 10005 ] |
| Status | Needs 1st Level Review [ 10005 ] | First Level Review in Progress [ 10301 ] |
| Status | First Level Review in Progress [ 10301 ] | Ready for Pull Request [ 10304 ] |
| Status | Ready for Pull Request [ 10304 ] | Pull Request Submitted [ 10101 ] |
| Status | Pull Request Submitted [ 10101 ] | Reviewing Pull Request [ 10303 ] |
| Status | Reviewing Pull Request [ 10303 ] | Merged Needs Testing [ 10002 ] |
| Status | Merged Needs Testing [ 10002 ] | Post-merge Testing In Progress [ 10003 ] |
| Resolution | Done [ 10000 ] | |
| Status | Post-merge Testing In Progress [ 10003 ] | Closed [ 6 ] |
| Epic Child |
|
| Epic Child | IGBF-3345 [ 22367 ] |
| Epic Child | IGBF-3366 [ 22389 ] |
| Epic Child | IGBF-3367 [ 22390 ] |
| Epic Child |
|
| Epic Child |
|
| Epic Child |
|
| Epic Child |
|
| Epic Child |
|
| Epic Child |
|
| Epic Child |
|
| Summary | Process Kelsey's Palanivelu Lab data | Run Nextflow with Kelsey's Palanivelu Lab 2023 data |
| Epic Name | Process Kelsey's Palanivelu Lab data | Process and deploy Palanivelu Lab data |
| Description |
For this task, process new data set from the Palanivelu lab. These data are from Kelsey
RR has downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please do the data processing in this directory: * /nobackup/tomato_genome/30-804059537-KP (kp for "Kelsey Pryze") Bitbucket repo: https://bitbucket.org/hotpollen/pistil-rna-seq To-do: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attach to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters. Attached: * Azenta (sequencing provider) data report, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
For this task, process new and old experimental data sets from the Palanivelu lab.
First process data from 2023. These data were generated by Kelse Pryze, graduate student in RP Lab. RR downloaded this 2023 data from Kelsey onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please process this data set in this directory: * /nobackup/tomato_genome/30-804059537-KP (kp for "Kelsey Pryze") To-do for each experimental data set: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attach to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters. Repo: * Bitbucket repo: https://bitbucket.org/hotpollen/pistil-rna-seq Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Comment |
[ *Next steps:*
* Generate scaled coverage graphs (Molly) * Generate junction files (Molly - ask Ann to explain how to do this) * Copy data to IGB Quickload host for visualization in IGB (Ann) * Create samples Excel spreadsheet needed to create annots.xml for Quickload site (Molly - first draft & Ann - final draft) ] |
| Assignee | Molly Davis [ molly ] |
| Attachment | Screen Shot 2023-02-09 at 9.40.46 AM.png [ 17678 ] |
| Attachment | KP_samples.csv [ 17686 ] |
| Attachment | KP_multiqc_report.html [ 17687 ] |
| Description |
For this task, process new and old experimental data sets from the Palanivelu lab.
First process data from 2023. These data were generated by Kelse Pryze, graduate student in RP Lab. RR downloaded this 2023 data from Kelsey onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please process this data set in this directory: * /nobackup/tomato_genome/30-804059537-KP (kp for "Kelsey Pryze") To-do for each experimental data set: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attach to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters. Repo: * Bitbucket repo: https://bitbucket.org/hotpollen/pistil-rna-seq Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ Notes: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to Azenta sequencing orders. The RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta, the sequencing company. The company then sent links to the resulting sequence data files to the RP lab, who downloaded them. We also got them and put them on the Charlotte HPC file system. We then proceeded to process them to generate files for visualization in IGB and "counts" files for processing using statistical analysis libraries developed for RNA-Seq data. The three data collections are: 1) 2022 Kelsie Pryze's unpollinated pistil experiment: We first obtained and processed these data in 2023. These data were generated by Kelse Pryze, graduate student in RP Lab. You can identify these samples by looking at their Azenta identifier. Also, in our various pipelines and Jira records, we have been referring to these data by the data we got them: the "KP 2023" dataset. These data are from an experiment done by KP in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers of four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the unpollinated pistils because they were all sequened at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to separate these to make it more clear that the sample generation was done separately from the unpollinated pistils. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. RR downloaded this 2023 data from Kelsey onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please process this data set in this directory: As of August 2023, we are storing the original data files in a folder named for the Azenta identifier: 30-804059537 : processed data The data are stored on * /nobackup/tomato_genome/30-804059537-KP (kp for "Kelsey Pryze") To-do for each experimental data set: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attach to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters. Repo: * Bitbucket repo: https://bitbucket.org/hotpollen/pistil-rna-seq Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Summary | Run Nextflow with Kelsey's Palanivelu Lab 2023 data | Process and deploy Palanivelu Lab data |
| Description |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ Notes: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to Azenta sequencing orders. The RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta, the sequencing company. The company then sent links to the resulting sequence data files to the RP lab, who downloaded them. We also got them and put them on the Charlotte HPC file system. We then proceeded to process them to generate files for visualization in IGB and "counts" files for processing using statistical analysis libraries developed for RNA-Seq data. The three data collections are: 1) 2022 Kelsie Pryze's unpollinated pistil experiment: We first obtained and processed these data in 2023. These data were generated by Kelse Pryze, graduate student in RP Lab. You can identify these samples by looking at their Azenta identifier. Also, in our various pipelines and Jira records, we have been referring to these data by the data we got them: the "KP 2023" dataset. These data are from an experiment done by KP in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers of four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the unpollinated pistils because they were all sequened at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to separate these to make it more clear that the sample generation was done separately from the unpollinated pistils. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. RR downloaded this 2023 data from Kelsey onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. Please process this data set in this directory: As of August 2023, we are storing the original data files in a folder named for the Azenta identifier: 30-804059537 : processed data The data are stored on * /nobackup/tomato_genome/30-804059537-KP (kp for "Kelsey Pryze") To-do for each experimental data set: * Run nf-core/rnaseq pipeline with SL5/2022 target genome assembly using "reverse" strandedness parameter. * Check the multi-qc report (attach to this ticket). Re-run processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters. Repo: * Bitbucket repo: https://bitbucket.org/hotpollen/pistil-rna-seq Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ Notes: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to Azenta sequencing orders. The RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta, the sequencing company. The company then sent links to the resulting sequence data files to the RP lab, who downloaded them. We also got them and put them on the Charlotte HPC file system. We then proceeded to process them to generate files for visualization in IGB and "counts" files for processing using statistical analysis libraries developed for RNA-Seq data. The three data collections are: 1) 2022 Kelsie Pryze's unpollinated pistil heat stress experiment: These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded this 2023 data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. We then began processing these data in spring of 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-804059537. Also, in our various pipelines and Jira records, we have been referring to these data by the data we started working with them: the "KP 2023" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2022. When we deploy these data to the genome browser for visualization, we will probably use a different name that better describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. 2) self-pollinated pistil and style heat stress experiment This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples **More information to be added** 3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. **More information to be added** To-do for each experimental data set: * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyiesusing "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Resolution | Done [ 10000 ] | |
| Status | Closed [ 6 ] | To-Do [ 10305 ] |
| Description |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ Notes: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to Azenta sequencing orders. The RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta, the sequencing company. The company then sent links to the resulting sequence data files to the RP lab, who downloaded them. We also got them and put them on the Charlotte HPC file system. We then proceeded to process them to generate files for visualization in IGB and "counts" files for processing using statistical analysis libraries developed for RNA-Seq data. The three data collections are: 1) 2022 Kelsie Pryze's unpollinated pistil heat stress experiment: These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded this 2023 data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/30-804059537-kelsie. We then began processing these data in spring of 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-804059537. Also, in our various pipelines and Jira records, we have been referring to these data by the data we started working with them: the "KP 2023" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2022. When we deploy these data to the genome browser for visualization, we will probably use a different name that better describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. 2) self-pollinated pistil and style heat stress experiment This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples **More information to be added** 3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. **More information to be added** To-do for each experimental data set: * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyiesusing "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ Notes: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to Azenta sequencing orders. The RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta, the sequencing company. The company then sent links to the resulting sequence data files to the RP lab, who downloaded them. We also got them and put them on the Charlotte HPC file system. We then proceeded to process them to generate files for visualization in IGB and "counts" files for processing using statistical analysis libraries developed for RNA-Seq data. The three data collections are: 1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536 These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The samples were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. 2) self-pollinated pistil and style heat stress experiment We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples **More information to be added** 3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. **More information to be added** To-do for each experimental data set: * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Description |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ Notes: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to Azenta sequencing orders. The RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta, the sequencing company. The company then sent links to the resulting sequence data files to the RP lab, who downloaded them. We also got them and put them on the Charlotte HPC file system. We then proceeded to process them to generate files for visualization in IGB and "counts" files for processing using statistical analysis libraries developed for RNA-Seq data. The three data collections are: 1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536 These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The samples were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. 2) self-pollinated pistil and style heat stress experiment We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples **More information to be added** 3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. **More information to be added** To-do for each experimental data set: * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ Notes: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to Azenta sequencing orders. The RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta, the sequencing company. The company then sent links to the resulting sequence data files to the RP lab, who downloaded them. We also got them and put them on the Charlotte HPC file system. We then proceeded to process them to generate files for visualization in IGB and "counts" files for processing using statistical analysis libraries developed for RNA-Seq data. The three data collections are: 1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536 These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The samples were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. 2) self-pollinated pistil and style heat stress experiment We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples 3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. To-do for each experimental data set: * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Description |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ Notes: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to Azenta sequencing orders. The RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta, the sequencing company. The company then sent links to the resulting sequence data files to the RP lab, who downloaded them. We also got them and put them on the Charlotte HPC file system. We then proceeded to process them to generate files for visualization in IGB and "counts" files for processing using statistical analysis libraries developed for RNA-Seq data. The three data collections are: 1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536 These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The samples were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. 2) self-pollinated pistil and style heat stress experiment We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples 3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. To-do for each experimental data set: * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ Notes: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to Azenta sequencing orders. The RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta, the sequencing company. The company then sent links to the resulting sequence data files to the RP lab, who downloaded them. We also got them and put them on the Charlotte HPC file system. We then proceeded to process them to generate files for visualization in IGB and "counts" files for processing using statistical analysis libraries developed for RNA-Seq data. The three data collections are: 1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536 These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The samples were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. 2) self-pollinated stigma+style heat stress experiment We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas * tissue type: dissected stigma and style tissue from self-pollinated flowers There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples 3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. The data from this experiment are stored on the UNC Charlotte cluster in: /projects/tomato_genome/rnaseq/ravi-tamaulipas The azenta code for this sequencing experiment was: ???? *To-do for each experimental data set:* * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Description |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ Notes: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to Azenta sequencing orders. The RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta, the sequencing company. The company then sent links to the resulting sequence data files to the RP lab, who downloaded them. We also got them and put them on the Charlotte HPC file system. We then proceeded to process them to generate files for visualization in IGB and "counts" files for processing using statistical analysis libraries developed for RNA-Seq data. The three data collections are: 1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536 These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The samples were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. 2) self-pollinated stigma+style heat stress experiment We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas * tissue type: dissected stigma and style tissue from self-pollinated flowers There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples 3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. The data from this experiment are stored on the UNC Charlotte cluster in: /projects/tomato_genome/rnaseq/ravi-tamaulipas The azenta code for this sequencing experiment was: ???? *To-do for each experimental data set:* * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ Notes: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to Azenta sequencing orders. The RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta, the sequencing company. The company then sent links to the resulting sequence data files to the RP lab, who downloaded them. We also got them and put them on the Charlotte HPC file system. We then proceeded to process them to generate files for visualization in IGB and "counts" files for processing using statistical analysis libraries developed for RNA-Seq data. The three data collections are: 1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536 These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The samples were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. 2) self-pollinated stigma+style heat stress experiment We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas * tissue type: dissected stigma and style tissue from self-pollinated flowers There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples 3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. The data from this experiment are stored on the UNC Charlotte cluster in: /projects/tomato_genome/rnaseq/ravi-tamaulipas Rob downloaded these data from the sequencing provider on or around December 15, 2021. (This is the date that Rob created a Google Doc describing the files available from the sequencing provider's data transfer ftp site.) The azenta code for this sequencing experiment was: 30-681594536 *To-do for each experimental data set:* * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Description |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ Notes: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to Azenta sequencing orders. The RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta, the sequencing company. The company then sent links to the resulting sequence data files to the RP lab, who downloaded them. We also got them and put them on the Charlotte HPC file system. We then proceeded to process them to generate files for visualization in IGB and "counts" files for processing using statistical analysis libraries developed for RNA-Seq data. The three data collections are: 1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536 These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The samples were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. 2) self-pollinated stigma+style heat stress experiment We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas * tissue type: dissected stigma and style tissue from self-pollinated flowers There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples 3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. The data from this experiment are stored on the UNC Charlotte cluster in: /projects/tomato_genome/rnaseq/ravi-tamaulipas Rob downloaded these data from the sequencing provider on or around December 15, 2021. (This is the date that Rob created a Google Doc describing the files available from the sequencing provider's data transfer ftp site.) The azenta code for this sequencing experiment was: 30-681594536 *To-do for each experimental data set:* * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ *About the data*: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to "batches" of RNA samples that were sent to GeneWiz/Azenta for library synthesis and subsequent sequencing. For two of these batches, the RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta (formerly GeneWiz), the sequencing company. For one of these batches, the so-called "library synthesis pilot," the RP lab synthesized the libraries themselves and then sent the libraries to Azenta. Once the sequencing data are complete, the company sent links to an ftp site containing the data files to the RP lab, who downloaded them or asked us to download them. We then obtained the sequence data and deployed them to the Charlotte HPC file system for the next steps - data processing, in which we generate files for visualization in IGB and, also, "counts" files for statistical analysis libraries developed for RNA-Seq data. The three data collections are: *1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536* These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The samples were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. 2) self-pollinated stigma+style heat stress experiment We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas * tissue type: dissected stigma and style tissue from self-pollinated flowers There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples 3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment, Azenta id 30-605730043 This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. The data from this experiment are stored on the UNC Charlotte cluster in: /projects/tomato_genome/rnaseq/ravi-tamaulipas Rob downloaded these data from the sequencing provider on or around December 15, 2021. (This is the date that Rob created a Google Doc describing the files available from the sequencing provider's data transfer ftp site.) Note: We need to confirm if that the sequences obtained from the unpollinated pistils were from the same experiment as (1) above. If yes, which "replicate" were they? This will influence how we present the data in IGB. *To-do for each experimental data set:* * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Description |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ *About the data*: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to "batches" of RNA samples that were sent to GeneWiz/Azenta for library synthesis and subsequent sequencing. For two of these batches, the RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta (formerly GeneWiz), the sequencing company. For one of these batches, the so-called "library synthesis pilot," the RP lab synthesized the libraries themselves and then sent the libraries to Azenta. Once the sequencing data are complete, the company sent links to an ftp site containing the data files to the RP lab, who downloaded them or asked us to download them. We then obtained the sequence data and deployed them to the Charlotte HPC file system for the next steps - data processing, in which we generate files for visualization in IGB and, also, "counts" files for statistical analysis libraries developed for RNA-Seq data. The three data collections are: *1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536* These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The samples were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. 2) self-pollinated stigma+style heat stress experiment We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas * tissue type: dissected stigma and style tissue from self-pollinated flowers There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples 3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment, Azenta id 30-605730043 This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. The data from this experiment are stored on the UNC Charlotte cluster in: /projects/tomato_genome/rnaseq/ravi-tamaulipas Rob downloaded these data from the sequencing provider on or around December 15, 2021. (This is the date that Rob created a Google Doc describing the files available from the sequencing provider's data transfer ftp site.) Note: We need to confirm if that the sequences obtained from the unpollinated pistils were from the same experiment as (1) above. If yes, which "replicate" were they? This will influence how we present the data in IGB. *To-do for each experimental data set:* * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ *About the data*: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to "batches" of RNA samples that were sent to GeneWiz/Azenta for library synthesis and subsequent sequencing. For two of these batches, the RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta (formerly GeneWiz), the sequencing company. For one of these batches, the so-called "library synthesis pilot," the RP lab synthesized the libraries themselves and then sent the libraries to Azenta. Once the sequencing data are complete, the company sent links to an ftp site containing the data files to the RP lab, who downloaded them or asked us to download them. We then obtained the sequence data and deployed them to the Charlotte HPC file system for the next steps - data processing, in which we generate files for visualization in IGB and, also, "counts" files for statistical analysis libraries developed for RNA-Seq data. The three data collections are: *1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536* These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The samples were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. *2) self-pollinated stigma+style heat stress experiment, Azenta id 30-804059537* We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas * tissue type: dissected stigma and style tissue from self-pollinated flowers There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples *3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment, Azenta id 30-605730043* This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. The data from this experiment are stored on the UNC Charlotte cluster in: /projects/tomato_genome/rnaseq/ravi-tamaulipas Rob downloaded these data from the sequencing provider on or around December 15, 2021. (This is the date that Rob created a Google Doc describing the files available from the sequencing provider's data transfer ftp site.) Note: We need to confirm if that the sequences obtained from the unpollinated pistils were from the same experiment as (1) above. If yes, which "replicate" were they? This will influence how we present the data in IGB. *To-do for each experimental data set:* * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Description |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ *About the data*: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to "batches" of RNA samples that were sent to GeneWiz/Azenta for library synthesis and subsequent sequencing. For two of these batches, the RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta (formerly GeneWiz), the sequencing company. For one of these batches, the so-called "library synthesis pilot," the RP lab synthesized the libraries themselves and then sent the libraries to Azenta. Once the sequencing data are complete, the company sent links to an ftp site containing the data files to the RP lab, who downloaded them or asked us to download them. We then obtained the sequence data and deployed them to the Charlotte HPC file system for the next steps - data processing, in which we generate files for visualization in IGB and, also, "counts" files for statistical analysis libraries developed for RNA-Seq data. The three data collections are: *1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536* These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The samples were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the sample generation was done separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. *2) self-pollinated stigma+style heat stress experiment, Azenta id 30-804059537* We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas * tissue type: dissected stigma and style tissue from self-pollinated flowers There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples *3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment, Azenta id 30-605730043* This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas. The data from this experiment are stored on the UNC Charlotte cluster in: /projects/tomato_genome/rnaseq/ravi-tamaulipas Rob downloaded these data from the sequencing provider on or around December 15, 2021. (This is the date that Rob created a Google Doc describing the files available from the sequencing provider's data transfer ftp site.) Note: We need to confirm if that the sequences obtained from the unpollinated pistils were from the same experiment as (1) above. If yes, which "replicate" were they? This will influence how we present the data in IGB. *To-do for each experimental data set:* * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblyies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset * Add the "counts" data files to the repository for statistical analysis Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
For this task,
* process new and old experimental data sets from the Ravi Palanivelu lab * confer with Palanivelu lab personnel to understand and document the samples * track code and key data files using this repository: https://bitbucket.org/hotpollen/pistil-rna-seq/src/main/ *About the data*: As of summer 2023, there are now three collections of sequencing data that we got from the Ravi Palanivelu lab. These collections correspond to "batches" of RNA samples that were sent to GeneWiz/Azenta for library synthesis and subsequent sequencing. For two of these batches, the RP lab created the biological material for samples, extracted RNA, and then sent the RNA boxes to Azenta (formerly GeneWiz), the sequencing company. For one of these batches, the so-called "library synthesis pilot," the RP lab synthesized the libraries themselves and then sent the libraries to Azenta. Once the sequencing data are complete, the company sent links to an ftp site containing the data files to the RP lab, who downloaded them or asked us to download them. We then obtained the sequence data and deployed them to the Charlotte HPC file system for the next steps - data processing, in which we generate files for visualization in IGB and, also, "counts" files for statistical analysis libraries developed for RNA-Seq data. The three data collections are: *1) 2021 Kelsie Pryze's unpollinated pistil heat stress experiment, azenta id: 30-681594536* These data are from an experiment done by Kelse Pryze in which she tested the effects of heat stress on un-pollinated tomato pistils dissected from emasculated flowers from four tomato varieties: Heinz, Malintka, Tamaulipas, and Nagcarlang. All sample types have three replicates per sample type represented in the sequencing data, except for Tamaulipas, which has two. KP provided a detailed description of exactly how the samples were generated. The biological material were created in 2021, in the summer and early spring. The data files also included three data files from a different experiment investigating the transcriptome of dissected, unpollinated ovary tissue. We processed these data alongside the fastq files from the un-pollinated pistils because they were all sequenced at the same time, in the same lot of RNAs sent to the sequencing provider. However, for visualization, we will probably want to present them in ways that will make it super clear that the biological material were created separately from the un-pollinated pistils. Rob Reid downloaded the data onto the UNCC cluster and saved it here: /projects/tomato_genome/rnaseq/ravi-2022-fullrun/30-681594536. We then began processing these data in 2023, using the high performance computing cluster at UNC Charlotte. You can identify these samples on our file system by looking for their Azenta identifier - 30-681594536. Also, in our various pipelines and Jira records, we have been referring to these data by the date we got them from the RP Lab: the "Ravi 2022" dataset. KP provided documentation describing these samples. We will place these files in a "Documentation" folder in the git repository. However, as you will see from the documentation in the repository, the samples themselves were generated during the summer and early spring of 2021. When we deploy these data to the genome browser for visualization, we will probably use a study name that describes the data and makes it easy for RP Lab personnel and others to recognize them in the browser or other settings. *2) self-pollinated stigma+style heat stress experiment, Azenta id 30-804059537* We have been referring to this experimental data set as "KP-2023", referring to the experimenter Kelsie Pryze and the date when we obtained the experimental data sequence files. The original sequence files are downloaded to this location on the UNC Charlotte cluster computing system: /projects/tomato_genome/rnaseq/30-804059537-kelsie This experiment included sample types testing two temperature conditions, three treatment durations, four varieties, and one tissue type. These were: * temperature conditions: 37 degrees C (heat stress) and 25 degrees C (control) * treatment durations: 0 hours (no heat stress applied), 3 hours, and 8 hours * four varieties: Heinz, Malintka, Nagcarlang, Tamaulipas * tissue type: dissected stigma and style tissue from self-pollinated flowers There were three replicates per sample type. The zero-hour samples however included three 25 degrees C samples and no 37 degrees C samples. Number of samples: (2 conditions * 4 varieties * 2 treatment durations * 3 replications) + (1 condition * 4 * 1 treatment duration (0 hours) * 3 replications ) = 60 samples *3) self-pollinated and unpollinated Tamaulipas library preparation pilot experiment, Azenta id 30-605730043* This experiment performed by Kelsie Pryze involved creating libraries for sequencing using RNAs from pollinated and upollinated samples from Tamaulipas plants. The data from this experiment are stored on the UNC Charlotte cluster in: /projects/tomato_genome/rnaseq/ravi-tamaulipas Rob downloaded these data from the sequencing provider on or around December 15, 2021. (This is the date that Rob created a Google Doc describing the files available from the sequencing provider's data transfer ftp site.) Note: We need to confirm if that the sequences obtained from the unpollinated pistils were from the same experiment as (1) above. If yes, which "replicate" were they? This will influence how we label the data in IGB. *To-do for each experimental data set:* * Run nf-core/rnaseq pipeline with both the SL5/2022 and SL4/2019 target genome assemblies using "reverse" strandedness parameter. * Check the multi-qc report. Re-run the processing as necessary. * Rename BAM files to not included "sorted" in the name. * Create scaled coverage graphs. * Create junction files. * Migrate data to an on-line location for IGB visualization. * Create annots.xml metadata file with visualization parameters for each dataset; add the data collection to the makeAnnotsXml.py script * Add the "counts" data files to the repository for statistical analysis * Add documentation for each sequence collection to the git repository * Perform data checking to catch any record-keeping errors that may have occurred Attached: * Azenta (sequencing provider) data report for KP 2023 data, with numbers of sequences produced * Quote from Azenta indicating strand-specific RNA-Seq, 2x150 bp paired end sequencing Contact: * Kelsey Pryze - kelseypryze@email.arizona.edu |
| Epic Child |
|
| Epic Child |
|
| Epic Child |
|
| Epic Child |
|