Details
-
Type:
Task
-
Status: Closed (View Workflow)
-
Priority:
Minor
-
Resolution: Done
-
Affects Version/s: None
-
Fix Version/s: None
-
Labels:
-
Story Points:2
-
Epic Link:
-
Sprint:Summer 1
Description
Situation: psl files (EST and mRNA) have some kind of off by one error. The listed end position is +1 compared to where the annotations actually end (positive strand alignments).
For example, if a mRNA on the positive strand says that it starts at 100 and ends at 150, in IGB the annotation will be drawn overlapping bases 100 - 149. If on the negative strand the IGB reported start is +1 to where the annotation is actually drawn. This indicates that the tEnd (https://genome.ucsc.edu/FAQ/FAQformat.html#format2) column is effectively +1 compared to what is actually drawn in IGB. Note that the annotations in IGB appear to be drawn in the correct location, i.e. intron exon boundaries line up correctly. The only issue is that the IGB reported start/end is off by one, depending on positive/negative strand.
Task: Investigate why this issue is occurring. If the issue can be fixed update this ticket.
Attachments
Issue Links
- relates to
-
IGBF-3330 Add mm39 (GRCm39) mouse genome to IGB
-
- Closed
-
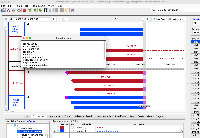
This issue is present in older psl files (M_musculus_Dec_2011) and on older versions of IGB (tested 9.0.2).