Details
-
Type:
Task
-
Status: Closed (View Workflow)
-
Priority:
Major
-
Resolution: Done
-
Affects Version/s: None
-
Fix Version/s: None
-
Labels:None
-
Story Points:1
-
Epic Link:
-
Sprint:Fall 1 - Sep 5
Description
Relates to error: IGBF-3420
Error:
Exception in thread "main" java.lang.RuntimeException: Postion is too high (more than 64792705)
at org.biojava.nbio.genome.parsers.twobit.TwoBitParser.setCurrentSequencePosition(TwoBitParser.java:191)
at org.biojava.nbio.genome.parsers.twobit.TwoBitParser.loadFragment(TwoBitParser.java:332)
at org.lorainelab.findjunctions.FindJunctions.main(FindJunctions.java:249)
Issue: Looked at the bam file which looks fine but there are reads that expand upon the chromosome. They have been soft clipped but find_junctions isn't taking that into consideration and won't process the file entirely to create the .FJ.bed.gz file.
Visualize the error in IGB.
Files used to visualize error:
Directory: /projects/tomato_genome/fnb/dataprocessing/30-804059537-KP/S_lycopersicum_Sep_2019/results/star_salmon
Visualization of the Junction Error in IGB:
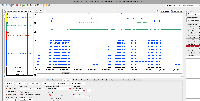
Issue was seen at end of chromosome 10 SL4. You can see in the red track that a lot of data is missing for the junction file, while in the green the junction file is normal and was processed correctly.
End of chromosome 10: