Details
-
Type:
Task
-
Status: Closed (View Workflow)
-
Priority:
Major
-
Resolution: Done
-
Affects Version/s: None
-
Fix Version/s: 10.1.0
-
Labels:None
-
Story Points:4
-
Epic Link:
-
Sprint:Fall 7, Fall 8, Spring 1, Spring 2, Spring 3, Spring 4
Description
Develop code to connect to this https://api.genome.ucsc.edu//getData/track?genome=hg38&track=ReMapDensity&chrom=chr1&start=1&end=248956 API and load Data for the Wiggle file type.
Attachments
Issue Links
- relates to
-
IGBF-3503 Implement UCSC REST API logic in IGB
-
- Closed
-
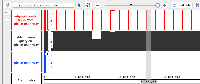
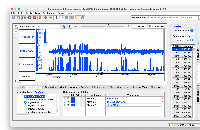
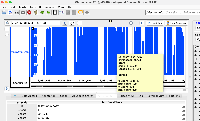
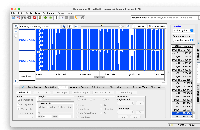
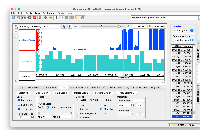
Implemented the code to load the wiggle format files (both bigWig and wig formats), and used existing GraphIntervalSym to load the syms. Testing is also done, working as expected. Didn't add the wigMaf file format as it's a bit different from the wiggle file formats: https://api.genome.ucsc.edu/getData/track?genome=hg38&track=multiz100way&chrom=chr1&start=8372&end=33172, Nowlan Freese Could you please check this format once and let me know whether it's okay to treat it as a wig format.
Code is available at branch: https://bitbucket.org/jaya-sravani/integrated-genome-browser/branch/IGBF-3501.
To test: