Details
-
Type:
Task
-
Status: Closed (View Workflow)
-
Priority:
Major
-
Resolution: Done
-
Affects Version/s: None
-
Fix Version/s: 10.1.0
-
Labels:None
-
Story Points:4
-
Epic Link:
-
Sprint:Fall 7, Fall 8, Spring 1, Spring 2, Spring 3, Spring 4
Description
Develop code to connect to this https://api.genome.ucsc.edu//getData/track?genome=hg38&track=ReMapDensity&chrom=chr1&start=1&end=248956 API and load Data for the Wiggle file type.
Attachments
Issue Links
- relates to
-
IGBF-3503 Implement UCSC REST API logic in IGB
-
- Closed
-
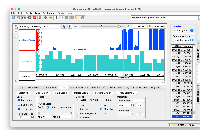
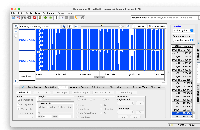
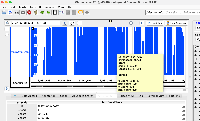
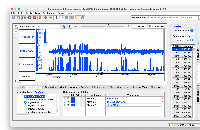
phastCons77way appearing correctly in IGB.
Closing ticket.