Details
-
Type:
Task
-
Status: Closed (View Workflow)
-
Priority:
Minor
-
Resolution: Done
-
Affects Version/s: None
-
Fix Version/s: None
-
Labels:None
-
Epic Link:
-
Sprint:Spring 4, Spring 5
Description
This task is more of a brainstorm based on the details that came out of Kelsey's committee meeting.
In her meeting, an advisor inquired about how Heinz is the reference genome and a sample in the experiment. It would make sense that Heinz will more easily align to reference genome compared to the other 3 TMH varieties.
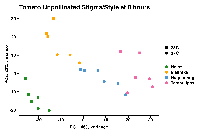
The PCA (attached) reflects this.
The goal would be to come up with an unbiased way to check MDS / PCA / PCoA where everything is not aligned to Heinz.
1 idea.
- We de novo assembly via trinity.
- Get a set of genes for each variety.
- Assign all of these genes to a SolyID via blast. Along with a gene expression count.
- Run these as a MDS / PCA and see the distribution.