Details
-
Type:
Bug
-
Status: To-Do (View Workflow)
-
Priority:
Blocker
-
Resolution: Unresolved
-
Labels:
Description
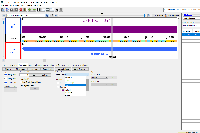
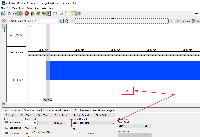
When I load an annotation file (gff3) using the Windows version of IGB, it does not show the option to show the gene name (gene=) from the gff3 file (as seen in the drop down menu). If I use this same file in the Linux version, it gives me the right option and a few more.
I would like to be able to label by gene name eg. "lysT" instead of this UNKNOWN_SYM65 id.
GFF file in this example region:
AL123456 EMBL gene 923803 923875 . - . ID=gene870;Name=lysT;gbkey=Gene;gene=lysT;gene_biotype=tRNA
AL123456 EMBL tRNA 923803 923875 . - . ID=rna13;Parent=gene870;Note=codon recognized: AAA - lysT - tRNA-Lys - anticodon ttt - length %3D 73;anticodon=(pos:complement(923840..923842));gbkey=tRNA;gene=lysT;product=tRNA-Lys
AL123456 EMBL exon 923803 923875 . - . ID=id50;Parent=rna13;Note=codon recognized: AAA - lysT - tRNA-Lys - anticodon ttt - length %3D 73;anticodon=(pos:complement(923840..923842));gbkey=tRNA;gene=lysT;product=tRNA-Lys
Reporter: Valdir Barth
E-mail: valdirbarth@gmail.com![]()
From: Mason Meyer <mmeyer20@uncc.edu>
To: Valdir Barth <valdirbarth@gmail.com>
Date: Thu, Nov 30, 2017 at 5:12 PM
Subject: Re: Windows version does not read gff file properly
Hello again Valdir,
I am glad you were able to find out what was causing the issue. I looked into this to see if other GFF3 files also seemed to have this issue and it seems to me that this issue may not be the norm. This must be something NCBI is doing to their GFF3 files, but IGB is actually reading the files as expected. Our team may be able to add some error-handling to files containing this additional line at the beginning, though. I will discuss this issue with the rest of the team and will track this in our issue-tracking system so we can plan work on this task.
Thank you for continuing to look into this issue. If you have any other questions or feedback for us, please be sure to let me know.
Thanks again,
Mason Meyer
IGB Support Specialist