Details
-
Type:
Improvement
-
Status: Closed (View Workflow)
-
Priority:
Major
-
Resolution: Done
-
Affects Version/s: None
-
Fix Version/s: 9.1.10 Major Release
-
Labels:None
-
Story Points:4
-
Epic Link:
-
Sprint:Fall 6 2021 Oct 25 - Nov 5, Fall 7 2021 Nov 8 - Nov 24, Fall 8 2021 Nov 29 - Dec 10, Fall 9 2021 Dec 13 - Dec 24, Spring 1 2022 Jan 3 - Jan 14
Description
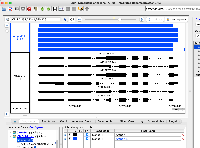
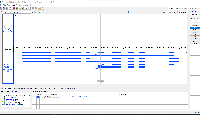
Situation: bigBed is an indexed binary format of bed files. However, in IGB, bigBed files are not appearing correctly. The bigBed gene annotations (see attached file) are not showing the introns/exons, but instead are drawing a single glyph for the entire gene.
Task: Fix IGB so that the bigBed files appear correctly.
See the UCSC guide on bigBed for more information on the bigBed file format.
Attachments
Issue Links
Activity
To reproduce the issue:
Open IGB
Open the Arabidopsis thaliana A_thaliana_Jun_2009 genome
Navigate to Chr1:2,257,253-2,260,326
Load the attached Araport11.bb file
Click Load Data and compare the Araport11.bb file and the Araport11 bed file found in IGB Quickload (should load by default)

If you are looking for example .as (autosql) files check out the following links:
https://github.com/ucscGenomeBrowser/kent/tree/e01be94b2df0b6b467170df7e304ed87493317bd/src/hg/lib
https://genome-source.gi.ucsc.edu/gitlist/kent.git/raw/master/src/hg/lib/