More thoughts:
A PCA needs as input a matrix of numbers. In our case, the matrix rows are genes, matrix columns are samples, and values are expression measurements per gene, per sample.
We can make PCA plots showing all the data (all samples) currently because we are using a common genome assembly as the alignment target. We have expression measurements for every gene, in every sample.
The gene models are from the reference Heinz assembly, in this case.
If we make RNA-Seq assemblies - a collection of transcript models - how will we get the matrix to use as input to PCA?
We would need a way to make the input matrix that wouldn't exacerbate bias arising from Heinz alignments working best against Heinz sequence.
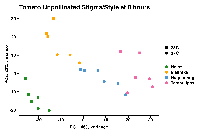
PCA plots give you a nice, visual overview of the distributions of variables in a data set. We interpret them by making note of 2-dimension relationships between points, where we use color and shape to indicate values of experimental variables - temperature, cultivar, age, etc. In our case, points cluster by cultivar. This means there are large differences in gene expression measurements associated with cultivar in the data set. These differences could arise from many causes, of which alignment quality *could* be one. Heinz sequences may align better to some genes than do the other varieties.
Is there a way that we can assess the degree to which alignment quality is affecting the results?
Has anyone looked at whether there are any signs in the data of such a bias? How do we do that?
Task
Minor
A more detailed plan laid out in this email:
I think I can churn something out in time. Let's lay out a plan first. I can start the pipeline and I have a student rotating with me who can help as well. I think we can have it done in time.
But if not, you can show the pipeline, lay out the justification and describe the expected end result.
It will also be a good idea to run the pipeline in Arizona afterwards. I share with you the code, you run it, we make it all publicly available for publication! You running it = repeatable.
Let me know if the following is a good strategy:
GOAL: To create gene counts for each plant variety using the same plant sequences and a de novo assembly approach to avoid HEINZ reference bias.
Also, I have Molly run and fine tune the pipeline, and she then sets up in BitBucket to make the code publicly available for others.